If you have ever used the R package lme4 to perform mixed-effect modeling you may have noticed the “Correlation of Fixed Effects” section at the bottom of the summary output. This article intends to shed some light on what this section means and how you might interpret it.
To begin, let’s simulate some data. Below we generate 30 observations for 20 subjects. The gl() function generates levels. In this case it generates 30 replications of 20 levels. The result is the numbers 1 - 20, each repeated 30 times. We name it id and can think of these as 20 subjects numbered 1 - 20. We then generate the sequence 1 - 30, 20 times, using the rep() function. This will serve as our predictor, or independent variable.
n <- 20 # subjects
k <- 30 # observations
id <- gl(n, k)
x <- rep(1:k, n)
Next we generate some “random noise” for each observation by drawing 20 * 30 = 600 samples from a Normal distribution with mean 0 and standard deviation 12. The value 12 is arbitrary. Feel free to try a different value. The function set.seed(1) allows you to sample the same data if you wish to follow along.
set.seed(1)
obs_noise <- rnorm(n * k, mean = 0, sd = 12)
We also need to generate noise for each subject to create data suitable for mixed-effect modeling. Again we sample from a Normal distribution with mean 0, but this time we arbitrarily set the standard deviation to 20. We also only sample 20 values instead of 600. That’s because the noise is specific to the subject instead of the observation. Run the set.seed(11) code if you want to sample the same data.
set.seed(11)
subj_noise <- rnorm(n, mean = 0, sd = 20)
Finally we generate a dependent variable, y, as a linear function of x using the formula y = 3 + 2*x. The fixed intercept is 3. The fixed slope is 2. We also include the noise. The code subj_noise[id] repeats the subject-specific noise 30 times for each subject and is added to the intercept. The observation-specific noise is added at the end. When done we place our variables in a data frame we name d.
y <- (3 + subj_noise[id]) + 2*x + obs_noise
d <- data.frame(id, y, x)
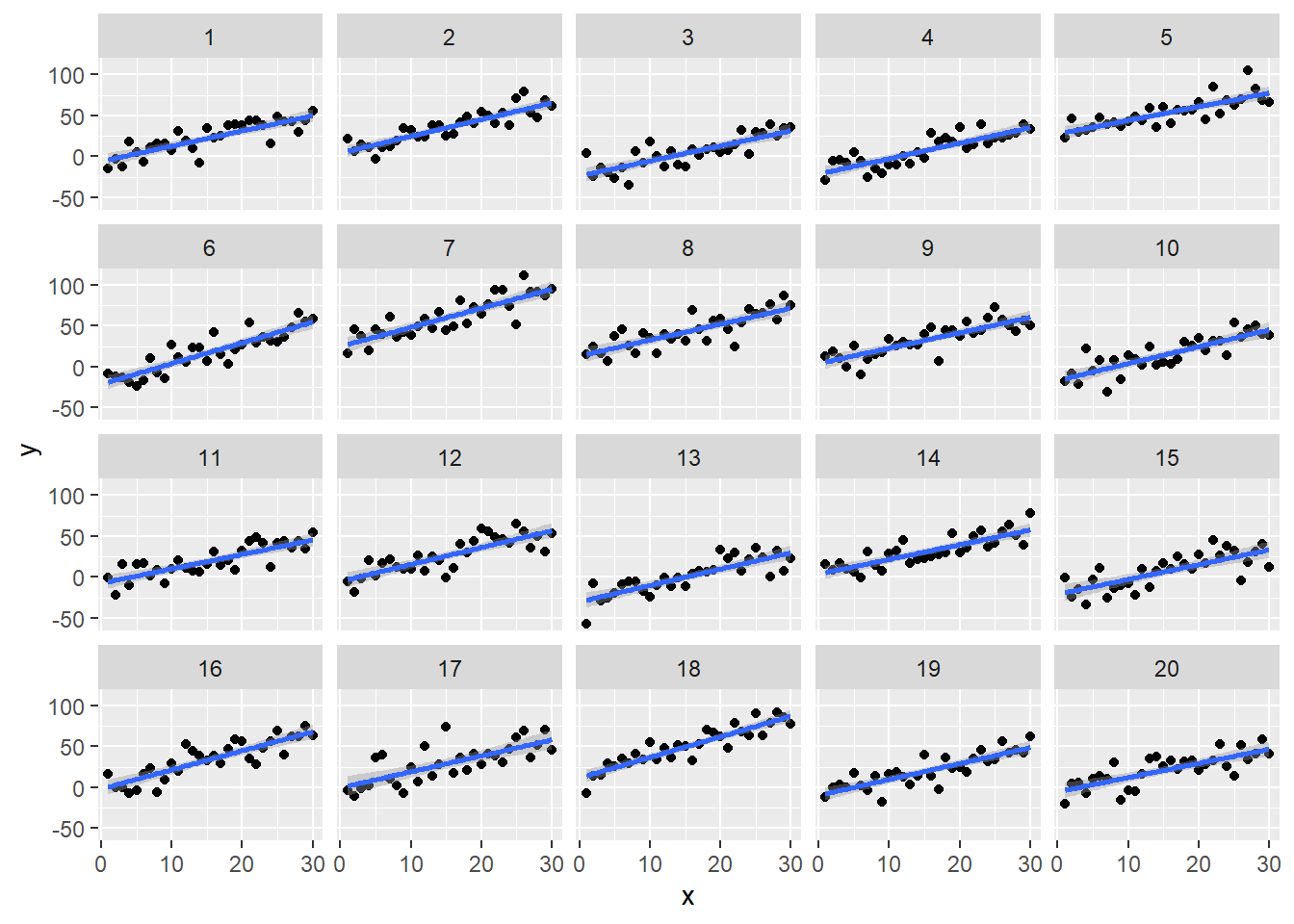
Let’s visualize our data using ggplot2. Notice the fitted line for each subject has about the same slope but shifts up or down for each subject. That’s how we simulated our data. Each subject has the same slope but a different intercept that changes according the subject-specific noise we sampled.
library(ggplot2)
ggplot(d) +
aes(x = x, y = y) +
geom_point() +
geom_smooth(method = "lm") +
facet_wrap(~id)
Now we use the lmer() function from the lme4 package to “work backwards” and try to recover the true values we used to generate the data. Notice we are fitting the “correct” model. We know the process we used to generate the data, so we know which model to specify.
library(lme4)
me1 <- lmer(y ~ x + (1|id), data = d)
summary(me1)
Linear mixed model fit by REML ['lmerMod']
Formula: y ~ x + (1 | id)
Data: d
REML criterion at convergence: 4771
Scaled residuals:
Min 1Q Median 3Q Max
-2.7171 -0.6417 -0.0160 0.6715 3.7493
Random effects:
Groups Name Variance Std.Dev.
id (Intercept) 267.6 16.36
Residual 146.1 12.09
Number of obs: 600, groups: id, 20
Fixed effects:
Estimate Std. Error t value
(Intercept) -3.843 3.795 -1.013
x 1.999 0.057 35.073
Correlation of Fixed Effects:
(Intr)
x -0.233
A quick review of the summary output shows the lmer() did quite well at recovering the data-generating values. Under the “Fixed Effects” section the slope of x is estimated as 1.99 which is very close to the true value of 2. The estimate of the intercept is not terribly good, but we added a fair amount of noise to it for each subject, so it’s not surprising lmer() struggled with it. Under the “Random Effects” section the standard deviations of the Normal distributions from which we sampled the noise are estimated as about 12.1 and 16.4, pretty close to the true values of 12 and 20.
And that brings us to the final section: Correlation of Fixed Effects. It shows that the intercept and x coefficients have a correlation of -0.233. What is that exactly? We certainly didn’t use correlated data to generate fixed effects. In fact we simply picked two numbers: 3 and 2. How can our fixed effects be correlated?
To help answer this, we’ll first point out the estimated standard errors for the intercept and x coefficients. The intercept standard error is 3.795. The x standard error is 0.057. These quantify uncertainty in our estimate. It may be helpful to think of them as “give or take” values. For example, our x coefficient is 1.99, give or take about 0.057. These standard errors are the square roots of the estimated variances. We can see the variances using the vcov function.
vcov(me1)
2 x 2 Matrix of class "dpoMatrix"
(Intercept) x
(Intercept) 14.4018199 -0.050363397
x -0.0503634 0.003249251
We can extract the diagonal values of the matrix using the diag() function and take the square roots to get the standard errors that are presented in the summary output. We use the base R pipe to send the output of one function to the input of the next.
vcov(me1) |> diag() |> sqrt()
[1] 3.79497298 0.05700221
If we look again at the output of the vcov() function we see a covariance estimate in the off-diagonal entry of the matrix: -0.0503634
vcov(me1)
2 x 2 Matrix of class "dpoMatrix"
(Intercept) x
(Intercept) 14.4018199 -0.050363397
x -0.0503634 0.003249251
So not only do we get an estimate of uncertainty for each coefficient, we get an estimate of how the coefficients vary together. Hence the name of the function, “vcov”, which is short for variance-covariance.
If we divide the covariance by the product of the standard errors, we get the correlation of fixed effects reported in the lmer() summary output.
m <- vcov(me1)
den <- m |> diag() |> sqrt() |> prod()
num <- m[2,1]
num/den
[1] -0.232817
In fact this is the formula for the familiar correlation coefficient.
\[\rho = \frac{\text{Cov}(X_1, X_2)}{\sigma_1\sigma_2}\]
Base R includes the convenient cov2cor() function to automatically make these calculations for us.
vcov(me1) |> cov2cor()
2 x 2 Matrix of class "dpoMatrix"
(Intercept) x
(Intercept) 1.000000 -0.232817
x -0.232817 1.000000
Now that we see where the correlation of fixed effects is coming from, what does it mean?
Imagine we replicate our analysis above many times, each time using a new random sample. We will get new estimates of fixed effects every time. The correlation of fixed effects gives us some sense of how those many fixed effect estimates might be associated. Let’s demonstrate this with a simulation.
To begin we use our model to simulate 1000 new sets of responses using the simulate() function. Recall the point of mixed-effect modeling is to estimate the data-generating process. Once we have our model we can then use it to generate data. (A good fitting model should generate data similar to the observed data.) The result is a data frame named “sim” with 1000 columns. Each column is a new vector of 600 responses.
sim <- simulate(me1, nsim = 1000)
After that we apply our lme4 model to each new set of responses. We set up our model as a function named “f” with the argument k. Then we apply that function to each column of the sim object using the lapply() function. The resulting object named “out” contains 1000 model fits with 1000 different sets of fixed effects. Depending on your computer, this may take anywhere from 10 to 30 seconds.
f <- function(k)lmer(k ~ x + (1|id), data = d)
out <- lapply(sim, f)
Once that finishes running we do a bit of data wrangling to extract the fixed effects into a matrix, calculate the correlation of the fixed effects, and create a scatter plot.
# extract fixed effects
f_out <- lapply(out, fixef)
# collapse list into matrix
f_df <- do.call(rbind, f_out)
cor(f_df[,1], f_df[,2])
[1] -0.2335858
The correlation of -0.234 is close to the Correlation of Fixed Effects reported in the original summary output, -0.233. We can also plot the simulated fixed effects. It appears higher coefficients of x are slightly associated with lower coefficients of the intercept.
plot(f_df)

Is this important to know? That’s up to you to decide. It’s another piece of information to help you understand your model. It just so happens this output is provided by default when using summary() on a model fitted with lmer(). If you prefer, you can suppress it by setting corr = FALSE. On the other hand, when calling summary() on a model fitted with lm() the correlation of fixed effects is not printed, but you can request it by setting corr = TRUE. The correlation of fixed effects is not unique to mixed-effect models.
We should note that examining the correlation of fixed effects is NOT the same as examining your model for collinearity. That involves the correlation of your predictors, not the model’s fixed effects.
R session details
The analysis was done using the R Statistical language (v4.5.1; R Core Team, 2025) on Windows 11 x64, using the packages lme4 (v1.1.37), Matrix (v1.7.3) and ggplot2 (v3.5.2).
References
- Bates D, Maechler M, Bolker B, Walker S (2015). Fitting Linear Mixed-Effects Models Using lme4. Journal of Statistical Software, 67(1), 1-48. https://www.jstatsoft.org/article/view/v067i01.
- R Core Team (2023). R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/.
Clay Ford
Statistical Research Consultant
University of Virginia Library
February 28, 2022
For questions or clarifications regarding this article, contact statlab@virginia.edu.
View the entire collection of UVA Library StatLab articles, or learn how to cite.
